{mmrm}: an Open Source R Package for Mixed Model Repeated Measures
(Statistical Engineering, Pharma Product Development Data Sciences, Roche)
2023-03-31
Acknowledgments
Authors:
- Brian Matthew Lang (MSD)
- Christian Stock (BI)
- Craig Gower-Page (Roche)
- Dan James (AstraZeneca)
- Daniel Sabanes Bove (Roche, lead)
- Doug Kelkhoff (Roche)
- Julia Dedic (Roche)
- Kevin Kunzmann (BI)
- Liming Li (Roche)
- Ya Wang (Gilead)
Acknowledgments
Thanks to:
- Ben Bolker (McMaster University)
- Davide Garolini (Roche)
- Dinakar Kulkarni (Roche)
- Gonzalo Duran Pacheco (Roche)
- Software Engineering working group (SWE WG)
Agenda
- Introduction and Overview of SWE WG
- Mixed Models for Repeated Measures - Why is it a Problem?
- MMRM Package
- Why this is not “yet another package”
- Comparing MMRM R Package to other Implementations
- Closing and Next Steps
Context: Software Engineering in Biostatistics
- Open-source software has gained increasing popularity in Biostatistics
- Pros: rapid uptake of novel statistical methods, unprecedented opportunities for collaboration
- Cons: variability in software quality
- Developing high-quality software is critical
Software Engineering working group (SWE WG)
- An official working group of the ASA Biopharmaceutical Section
- Formed in August 2022
- Cross-industry collaboration (>35 members from >25 organizations)
- Home page at rconsortium.github.io/asa-biop-swe-wg
- Deal with the issues of statistical quality assurances for R packages and create high quality statistical software
Why do we need a package for MMRM?
- Mixed Models for Repeated Measures (MMRM) is a popular choice for analyzing longitudinal continuous outcomes in randomized clinical trials
- No great R Package - initially thought that the MMRM problem was solved by using a combination of
lme4andlmerTest
- Learned that this approach failed on large data sets (slow, did not converge)
nlmedoes not give Satterthwaite adjusted degrees of freedom, has convergence issues, and withemmeansit is only approximate
Before creating a new package
- First try to improve existing package
- Here we tried to extend
glmmTMBto calculate Satterthwaite adjusted degrees of freedom - But it did not work
glmmTMBis always using a random effects representation, we cannot have a real unstructured model (uses \(\sigma = \varepsilon > 0\) trick)
- Here we tried to extend
- Think about long term maintenance and responsibility
Idea with some Details
- We only want to fit a fixed effects model with a structured covariance matrix for each subject
- The idea is then to use the Template Model Builder (
TMB) directly - as it is also underlyingglmmTMB- but code the exact model we want - We do this by implementing the log-likelihood in
C++using theTMBprovided libraries
Advantages of TMB
- Fast
C++framework for defining objective functions (Rcppwould have been alternative interface) - Automatic differentiation of the log-likelihood as a function of the variance parameters
- Syntactic sugars to allow simple matrix calculations or operations like R
- We get the gradient and Hessian exactly and without additional coding
- This can be used from the R side with the
TMBinterface and plugged into optimizers
Why it’s not just another package
- Ongoing maintenance and support from the pharmaceutical industry
- 5 companies being involved in the development, on track to become standard package
- Development using best practices as show case for high quality package
- Thorough unit and integration tests (also comparing with SAS results) to ensure accurate results
Features of {mmrm}
- Linear model for dependent observations within independent subjects
- Covariance structures for the dependent observations:
- Unstructured, Toeplitz, AR1, compound symmetry, ante-dependence, spatial exponential
- Allows group specific covariance estimates and weights
- REML or ML estimation, using multiple optimizers if needed
emmeansinterface for least square means- Satterthwaite adjusted degrees of freedom
- Kenward-Roger adjusted degrees of freedom
- Robust sandwich estimator for covariance
Comparing {mmrm} and SAS
To run an MMRM model in SAS it is recommended to use either the PROC MIXED or PROC GLM procedures.
Less model assumptions are applied in
PROC MIXED, primarily how one treats missingness.We will compares the
PROC MIXEDprocedure to the{mmrm}package in the following attributes:
- Documentation
- Unit Testing
- Covariance structures
- Covariance Estimators
- Degrees of Freedom
- Contrasts
Documentation
Both languages have online documentation of the technical details of the estimation and degrees of freedom methods and the different covariance structures available.
{mmrm}
PROC MIXED
Unit Testing
- Unit tests in
{mmrm}are transparent, compared toPROC MIXED. - Uses the
testthatframework withcovrto communicate the testing coverage. - Unit tests can be found in the GitHub repository under ./tests.
Note
The integration tests in {mmrm} are set to a tolerance of 10e^-3 when compared to SAS outputs.
Covariance structures
- SAS has 23 non-spatial covariance structures, while mmrm has 10.
- 9 structures intersect with SAS
- Ante-dependence (homogeneous) is only in the mmrm package.
- SAS has 14 spatial covariance structures compared to the spatial exponential one available in mmrm.
Tip
For users that need more structures, {mmrm} is easily extensible - let us know via feature requests in the GitHub repository.

Covariance structures details
| Covariance structures | {mmrm} |
PROC MIXED |
|---|---|---|
| Unstructured (Unweighted/Weighted) | X/X | X/X |
| Toeplitz (hetero/homo) | X/X | X/X |
| Compound symmetry (hetero/homo) | X/X | X/X |
| Auto-regressive (hetero/homo) | X/X | X/X |
| Ante-dependence (hetero/homo) | X/X | X |
| Spatial exponential | X | X |
Covariance Estimators
| Covariance estimators | {mmrm} |
PROC MIXED |
|---|---|---|
| Asymptotic | X | X |
| Kenward-Roger | X | X |
| Empirical | X | X |
| Jackknife | X | |
| Bias-Reduced Linearization | X |
Degrees of Freedom Methods
| Method | {mmrm} |
PROC MIXED |
|---|---|---|
| Contain | X* | X |
| Between/Within | X* | X |
| Residual | X* | X |
| Satterthwaite | X | X |
| Kenward-Roger | X | X |
| Kenward-Roger (Linear)** | X | X |
*Available through emmeans, and ongoing work in mmrm.
**This is not identical to the KR(Lin) in PROC MIXED.
Contrasts/LSMEANS
Contrasts and least square (LS) means estimates are available in mmrm using
mmrm::df_1d,mmrm::df_md- S3 method that is compatible with
emmeans - LS means difference can be produced through emmeans (
pairsmethod) - Degrees of freedom method is passed from
mmrmtoemmeans - By default
PROC MIXEDandmmrmdo not adjust for multiplicity, whereasemmeansdoes.
Note
These are comparable to the LSMEANS statement in PROC MIXED.
Benchmarking with other R packages
Other R packages, like nlme, lme4 and glmmTMB can support MMRM analysis. However:
- Covariance structure supported is limited
- Computaional efficiency is not satisfying
- Convergence issues(especially for
lme4) - Need other(multiple) packages to support
SatterthwaiteandKenward-Rogerdegrees of freedom, or Sandwich covariance estimator.
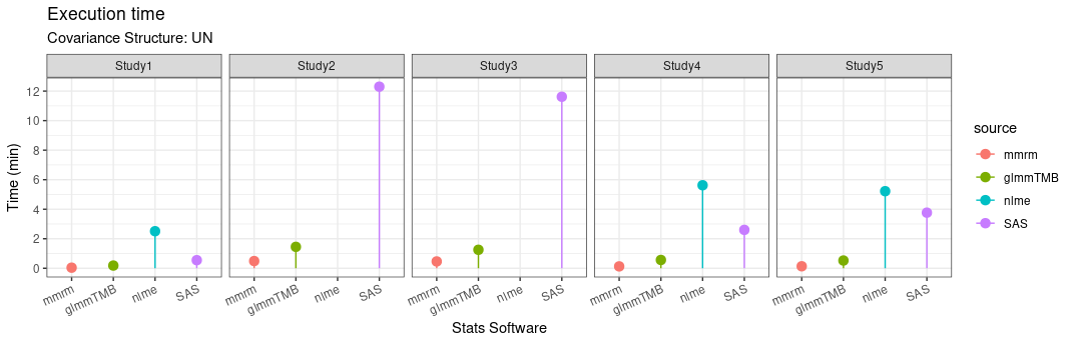
Computational Efficiency
mmrm not only supports multiple covariance structure, it also has good efficiency (due to fast implementations in C++)
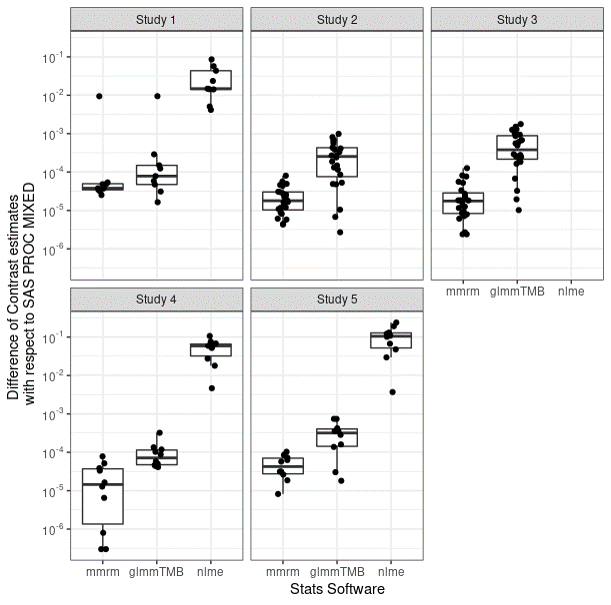
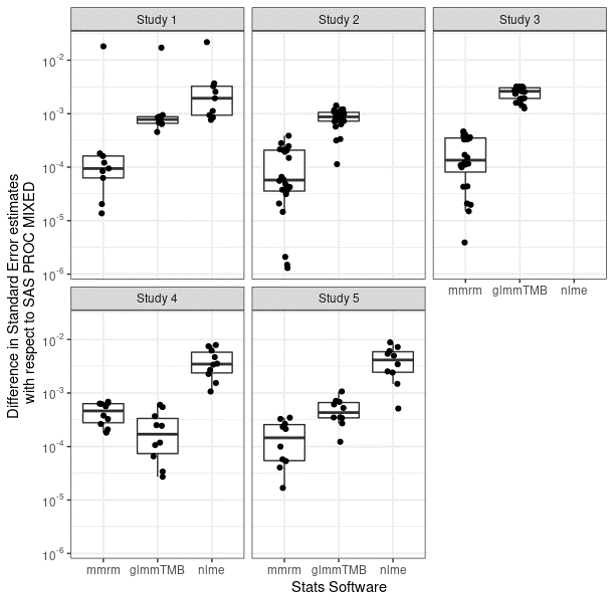
Differences with SAS
{mmrm} has small difference from SAS
Getting started
mmrmis on CRAN - use this as a starting point:
- Visit openpharma.github.io/mmrm for detailed docs including vignettes
- Consider tern.mmrm for high-level clinical reporting interface, incl. standard tables and graphs
Outlook
- We still have major features on our backlog:
- Type II and type III ANOVA tests
- Working on comparison to other implementations
- Parameter estimates, computation time, convergence, …
- Please let us know what is missing in
mmrmfor you! And try it out :-)
Thank you! Questions?


